The Broad Institute and MIT scientists who first harnessed CRISPR for mammalian genome editing have engineered a new molecular system for efficiently editing RNA in human cells. RNA editing, which can alter gene products without making changes to the genome, has profound potential as a tool for both research and disease treatment.
In a paper published today in Science, senior author Feng Zhang and his team describe the new CRISPR-based system, called RNA Editing for Programmable A to I Replacement, or “REPAIR.” The system can change single RNA nucleotides in mammalian cells in a programmable and precise fashion. REPAIR has the ability to reverse disease-causing mutations at the RNA level, as well as other potential therapeutic and basic science applications.
“The ability to correct disease-causing mutations is one of the primary goals of genome editing,” says Zhang, a core institute member of the Broad Institute, an investigator at the McGovern Institute, and the James and Patricia Poitras ’63 Professor in Neuroscience and associate professor in the departments of Brain and Cognitive Sciences and Biological Engineering at MIT. “So far, we’ve gotten very good at inactivating genes, but actually recovering lost protein function is much more challenging. This new ability to edit RNA opens up more potential opportunities to recover that function and treat many diseases, in almost any kind of cell.”
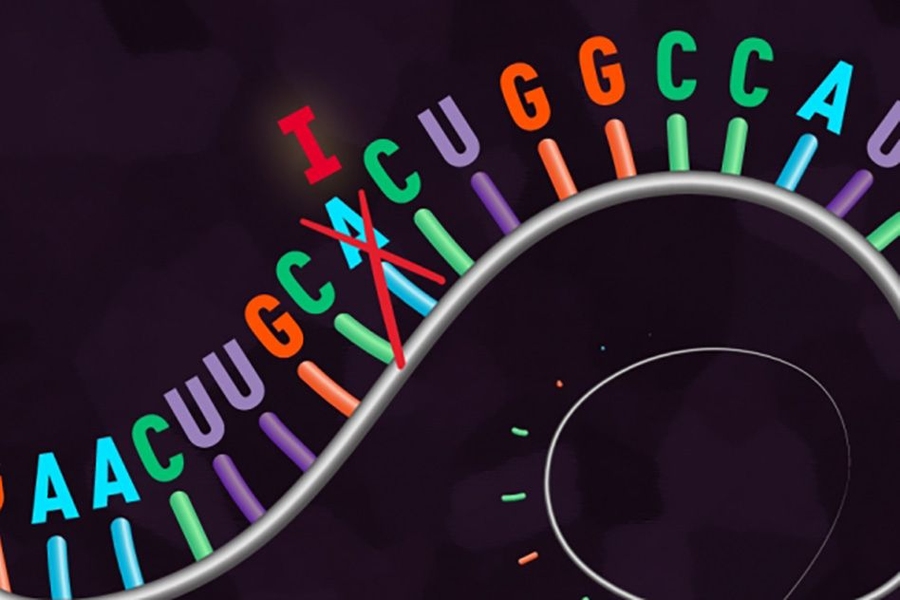
REPAIR has the ability to target individual RNA letters, or nucleosides, switching adenosines to inosines (read as guanosines by the cell). These letters are involved in single-base changes known to regularly cause disease in humans. In human disease, a mutation from G to A is extremely common; these alterations have been implicated in, for example, cases of focal epilepsy, Duchenne muscular dystrophy, and Parkinson’s disease. REPAIR has the ability to reverse the impact of any pathogenic G-to-A mutation regardless of its surrounding nucleotide sequence, with the potential to operate in any cell type.
Unlike the permanent changes to the genome required for DNA editing, RNA editing offers a safer, more flexible way to make corrections in the cell. “REPAIR can fix mutations without tampering with the genome, and because RNA naturally degrades, it’s a potentially reversible fix,” explains co-first author David Cox, a graduate student in Zhang’s lab.
To create REPAIR, the researchers systematically profiled the CRISPR-Cas13 enzyme family for potential “editor” candidates (unlike Cas9, the Cas13 proteins target and cut RNA). They selected an enzyme from Prevotella bacteria, called PspCas13b, which was the most effective at inactivating RNA. The team engineered a deactivated variant of PspCas13b that still binds to specific stretches of RNA but lacks its “scissor-like” activity, and fused it to a protein called ADAR2, which changes the letters A to I in RNA transcripts.
In REPAIR, the deactivated Cas13b enzyme seeks out a target sequence of RNA, and the ADAR2 element performs the base conversion without cutting the transcript or relying on any of the cell’s native machinery.
The team further modified the editing system to improve its specificity, reducing detectable off-target edits from 18,385 to only 20 in the whole transcriptome. The upgraded incarnation, REPAIRv2, consistently achieved the desired edit in 20 to 40 percent — and up to 51 percent — of a targeted RNA without signs of significant off-target activity. “The success we had engineering this system is encouraging, and there are clear signs REPAIRv2 can be evolved even further for more robust activity while still maintaining specificity,” says Omar Abudayyeh, co-first author and a graduate student in Zhang’s lab. Cox and Abudayyeh are both students in the Harvard-MIT Program in Health Sciences and Technology.
To demonstrate REPAIR’s therapeutic potential, the team synthesized the pathogenic mutations that cause Fanconi anemia and X-linked nephrogenic diabetes insipidus, introduced them into human cells, and successfully corrected these mutations at the RNA level. To push the therapeutic prospects further, the team plans to improve REPAIRv2’s efficiency and to package it into a delivery system appropriate for introducing REPAIRv2 into specific tissues in animal models.
The researchers are also working on additional tools for other types of nucleotide conversions. “There’s immense natural diversity in these enzymes,” says co-first author Jonathan Gootenberg, a graduate student in both Zhang’s lab and the lab of Broad core institute member Aviv Regev. “We’re always looking to harness the power of nature to carry out these changes.”
Zhang, along with the Broad Institute and MIT, plans to share the REPAIR system widely. As with earlier CRISPR tools, the groups will make this technology freely available for academic research via the Zhang lab’s page on the plasmid-sharing website Addgene, through which the Zhang lab has already shared reagents more than 42,000 times with researchers at more than 2,200 labs in 61 countries, accelerating research around the world.
This research was funded, in part, by the National Institutes of Health and the Poitras Center for Affective Disorders Research.