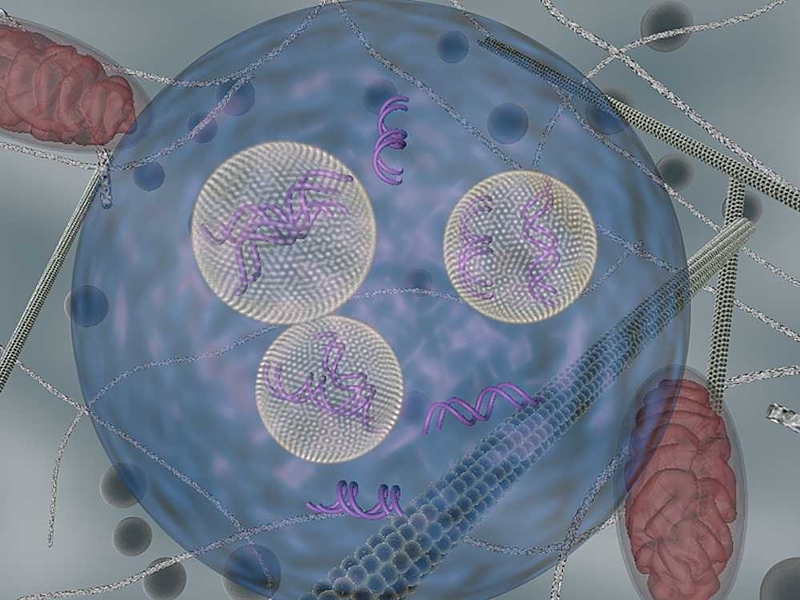
A new study from MIT sheds light on the nanoparticles’ fate and suggests new ways to maximize delivery of the RNA strands they are carrying, known as short interfering RNA (siRNA).
“We’ve been able to develop nanoparticles that can deliver payloads into cells, but we didn’t really understand how they do it,” says Daniel Anderson, the Samuel Goldblith Associate Professor of Chemical Engineering at MIT. “Once you know how it works, there’s potential that you can tinker with the system and make it work better.”
Anderson, a member of MIT’s Koch Institute for Integrative Cancer Research and MIT’s Institute for Medical Engineering and Science, is the leader of a research team that set out to examine how the nanoparticles and their drug payloads are processed at a cellular and subcellular level. Their findings appear in the June 23 issue of Nature Biotechnology. Robert Langer, the David H. Koch Institute Professor at MIT, is also an author of the paper.
One RNA-delivery approach that has shown particular promise is packaging the strands with a lipidlike material; similar particles are now in clinical development for liver cancer and other diseases.
Through a process called RNA interference, siRNA targets messenger RNA (mRNA), which carries genetic instructions from a cell’s DNA to the rest of the cell. When siRNA binds to mRNA, the message carried by that mRNA is destroyed. Exploiting that process could allow scientists to turn off genes that allow cancer cells to grow unchecked.
Scientists already knew that siRNA-carrying nanoparticles enter cells through a process, called endocytosis, by which cells engulf large molecules. The MIT team found that once the nanoparticles enter cells they become trapped in bubbles known as endocytic vesicles. This prevents most of the siRNA from reaching its target mRNA, which is located in the cell’s cytosol (the main body of the cell).
This happens even with the most effective siRNA delivery materials, suggesting that there is a lot of room to improve the delivery rate, Anderson says.
“We believe that these particles can be made more efficient. They’re already very efficient, to the point where micrograms of drug per kilogram of animal can work, but these types of studies give us clues as to how to improve performance,” Anderson says.
Molecular traffic jam
The researchers found that once cells absorb the lipid-RNA nanoparticles, they are broken down within about an hour and excreted from the cells.
They also identified a protein called Niemann Pick type C1 (NPC1) as one of the major factors in the nanoparticle-recycling process. Without this protein, the particles could not be excreted from the cells, giving the siRNA more time to reach its targets. “In the absence of the NPC1, there’s a traffic jam, and siRNA gets more time to escape from that traffic jam because there is a backlog,” says Gaurav Sahay, an MIT postdoc and lead author of the Nature Biotechnology paper.
In studies of cells grown in the lab without NPC1, the researchers found that the level of gene silencing achieved with RNA interference was 10 to 15 times greater than that in normal cells.
Lack of NPC1 also causes a rare lysosomal storage disorder that is usually fatal in childhood. The findings suggest that patients with this disorder might benefit greatly from potential RNA interference therapy delivered by this type of nanoparticle, the researchers say. They are now planning to study the effects of knocking out the NPC1 gene on siRNA delivery in animals, with an eye toward testing possible siRNA treatments for the disorder.
The researchers are also looking for other factors involved in nanoparticle recycling that could make good targets for possibly slowing down or blocking the recycling process, which they believe could help make RNA interference drugs much more potent. Possible ways to do that could include giving a drug that interferes with nanoparticle recycling, or creating nanoparticle materials that can more effectively evade the recycling process.
“This paper describes a new and very important way to improve the potency of siRNA delivery systems by inhibiting proteins that recycle imported material back out of the cell,” says Pieter Cullis, a professor of biochemistry and molecular biology at the University of British Columbia who was not part of the research team. “It is possible that this approach will give rise to the order-of-magnitude improvements in potency required for siRNA-based therapeutics to be more generally effective agents to treat disease.”
The research was funded by Alnylam Pharmaceuticals and the National Heart, Lung, and Blood Institute.