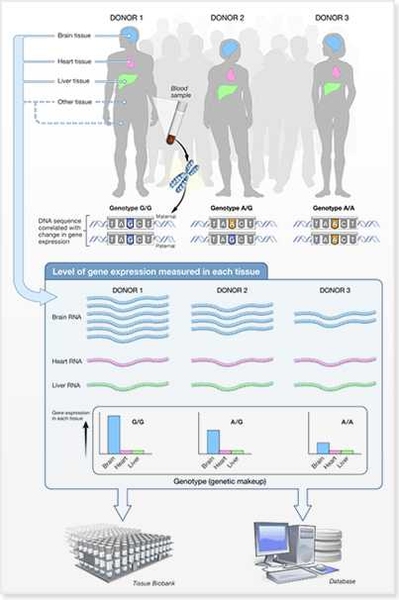
The National Institutes of Health have awarded eight grants as part of a new phase of the Genotype-Tissue Expression (GTEx) project to study the role that genomic variation plays in modulating gene expression and, ultimately, human disease. This new phase enhances GTEx to measure, in addition to gene expression levels, a diversity of molecular phenotypes that may ultimately influence human disease. The new grants explore inter-individual variation in DNA accessibility, protein levels, telomere lengths, and DNA modifications from diverse tissue samples whose collection began in 2010 across hundreds of individuals.
Manolis Kellis, professor of computer science at MIT, associate member of the Broad Institute of MIT and Harvard, and head of the MIT Computational Biology Group at the Computer Science and Artificial Intelligence Laboratory, will lead the MIT project. The goal is to measure changes in gene regulatory element activity in order to bridge the gap between genetic variation, gene expression, and human disease, by directly measuring the consequences of non-coding variants on gene regulation even before they affect gene expression levels.
Specifically, Kellis and his collaborators Alexander Meissner and Brad Bernstein from Harvard will work to characterize the epigenomic effects of genetic variation in nine peripheral tissues with roles in diabetes, heart disease, and cancer. The group will take an epigenomics approach that they previously used to study the regulatory circuitry of diverse human cell lines and tissues, and apply it to study how these elements vary across individuals.
“Genetic variants do not act in isolation, and their effects on organismal phenotypes are filtered through the complex circuitry of the cell”, explains Kellis. “For example, our risk for diabetes or cancer is an aggregate consequence of thousands of non-coding variants, acting in multiple tissues, affecting diverse genes, and ultimately influencing cellular and organismal functions. “
“This makes it very difficult to recognize the molecular basis of human disease, because our measurements are too far from the molecular consequences of genetic variants. This is why our project is focused on measuring directly the activity of hundreds of thousands of regulatory regions in each individual and each tissue.”
This strategy is based on the recognition that most genetic variants linked to disease don’t code for proteins, but instead have subtle gene regulatory roles, such as altering gene activity levels, or affecting the chemical modifications — epigenomic marks — made to DNA that influence which genes are active in which cells. The resulting information will be widely disseminated through the GTEx community portal, and used to guide the regulatory models developed by Kellis and the broader computational biology community, to guide their search for disease-causing mutations.
Kellis joined MIT as an undergraduate student in 1995, and earned his PhD in in 2003, earning the MIT Sprowls Award for the best thesis in computer science for his work on comparative genomics. His lab seeks to build a systematic understanding of the human genome, through computational integration of large-scale functional genomics, epigenomics, and comparative genomics. His group has developed a series of computational lenses for understanding genomes through their evolutionary signatures and epigenomic modifications to recognize functional elements in diverse cell types, and through activity-based regulatory signatures to link regulatory regions to their target genes.
Kellis has earned the U.S. Presidential Early Career Award in Science and Engineering (PECASE), the Alfred Sloan Foundation award, and the National Science Foundation CAREER award. His group has contributed to and helped lead many large-scale computational and experimental collaborative projects with the goal of understanding the human genome, and how genetic mutations contribute to complex traits and human disease.
The NIH Common Fund, which supports the GTEx, encourages collaboration and supports a series of high impact, trans-NIH programs. Common Fund programs are designed to pursue major opportunities and gaps in biomedical research that no single NIH Institute could tackle alone. Additional information about the NIH Common Fund can be found at http://commonfund.nih.gov.