A new study from MIT finds that two strains of bacteria that are each resistant to one antibiotic can protect each other in an environment containing both drugs.
The findings demonstrate that mutualism, a phenomenon in which different species benefit from their interactions with each other, can help bacteria form drug-resistant communities. This is the first experimental demonstration in microbes of a type of mutualism known as cross-protection, which is more commonly seen in larger animals.
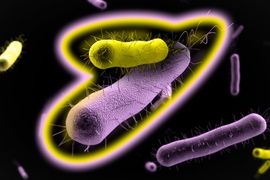
The researchers focused on two strains of E. coli, one resistant to ampicillin and the other resistant to chloramphenicol. These bacteria and many others defend themselves from antibiotics by producing enzymes that break down the antibiotics. As a side effect, this also protects cells that don’t produce those enzymes, by removing the antibiotic from the environment.
“Any time that you’re breaking down an antibiotic, there’s this potential for cross-protection,” says Jeff Gore, the Latham Family Career Development Associate Professor of Physics and the senior author of the study, which appears in the Proceedings of the National Academy of Sciences the week of May 16.
The MIT team found that, indeed, both strains could survive in an environment where both antibiotics were present, even though each was only resistant to one of the drugs. This type of situation is likely also found in the natural world, especially in soil where many strains of bacteria live together.
“Each of them is making different toxins and each of them is resistant to different toxins,” Gore says. “A lot of antibiotics are produced by microbes as part of the combat that is taking place between microorganisms in the soil.”
Gore and co-first authors Eugene Yurtsev and Arolyn Conwill, both MIT graduate students, also found that the populations of the two strains oscillate over time. Population oscillations are common in predator-prey interactions but rare in mutualistic interactions such as the cross-protection seen in this study.
Throughout their experiments, the researchers diluted the bacterial population each day by transferring about 1 percent of the population to a new test tube, to which new antibiotics were added. They found that while the total size of the bacterial population remained about the same, there were large oscillations in the relative percentages of each strain, which varied by nearly 1,000 percent over a period of about three days.
For example, if the ampicillin-resistant strain was more abundant in the beginning of a cycle, it rapidly deactivated ampicillin in the environment, allowing the chloramphenicol-resistant strain to begin growing. The ampicillin-resistant strain only began growing once the other strain had expanded enough to deactivate most of the chloramphenicol, at which point the chloramphenicol-resistant strain had already overtaken the ampicillin-resistant strain.
“The mutualism exhibits oscillations because the strain that is more abundant at the beginning of a growth cycle might end up less abundant at the end of that cycle,” Gore says.
At lower antibiotic concentrations, the bacterial population can survive in this oscillating pattern indefinitely, but at higher drug concentrations, the oscillations destabilize the population, and it eventually collapses.
Gore suspects that similar population oscillations may also be seen in natural environments such as the human gut, as bacteria exit the body along with bowel movements, or in soil as bacteria are washed away by rainfall.
Erdal Toprak, an assistant professor of systems biology at the University of Texas Southwestern Medical Center, says the study is an elegant demonstration of a seemingly simple interaction that may also be occurring in more complicated ecological systems.
“There may be many examples of this in the gut microbiome, and the tools developed in this study should be helpful to learn about more complex populations such as the gut microbiome or soil bacteria,” says Toprak, who was not involved in the study.
Gore’s lab is now looking at this type of mutualism in bacteria living in the gut of the worm C. elegans. The researchers are also studying how these types of population oscillations can become synchronized over large geographic areas, and how migration between populations influences this synchronization.