An MIT biophysicist's revelation of how bacteria clump as a defense mechanism may one day lead to drugs designed to make the bacteria act in ways that would allow us to get rid of them more easily.
Through exploring how Escherichia coli "remember" danger signals in their environment and interact with fellow organisms, Alexander van Oudenaarden, assistant professor of physics, describes in this week's Proceedings of the National Academy of Sciences the mechanism through which E. coli form clumps when they are threatened.
Although most strains of E. coli are harmless and live in the intestines of healthy humans and animals, a certain strain produces a powerful toxin and is an emerging cause of foodborne illness, causing an estimated 73,000 cases of infection and 61 deaths in the United States each year. Researchers also could extrapolate the behavior of these microbes to more harmful species such as cholera.
Van Oudenaarden collaborated on the work with MIT physics graduate student Nikhil Mittal; Elena Budrene-Kac, a research associate in the MIT Materials Processing Center; and Michael P. Brenner of Harvard's Division of Engineering and Applied Sciences.
Scientists knew E. coli, which move with the help of helical, thread-like tails called flagella, formed clusters when threatened, but they didn't know how these tiny cells organized themselves or how the cell community keeps these clusters stable.
Through experiments, theoretical modeling and computation, van Oudenaarden's group looks at how cells use genetic and biochemical networks to make smart decisions that enable them to survive. He works with bacteria and yeast--organisms that are biologically well defined and whose genomes are completely sequenced.
In the E. coli experiment, van Oudenaarden's group observed the effects of chemical signals on individual cells. A chemical communication system between bacterial cells tells each cell to forget its individual status and clump together with other cells into aggregates when stressed. These multicellular aggregates can then resist a toxic onslaught from chemicals such as antibiotics.
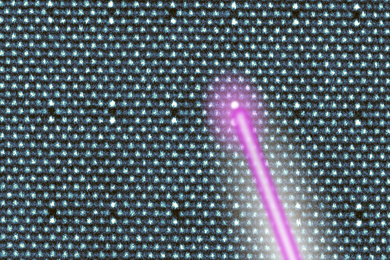
To look at individual organisms in a big community of millions of cells, van Oudenaarden labeled a handful of cells with a green fluorescent protein and then tracked them with fluorescent microscopy.
E. coli can move two ways. When the flagella rotate together counterclockwise, they move the organism straight ahead like a little propellor. When the flagella reverse from counterclockwise to clockwise, they become separate and disorganized, and the organism tumbles. After each tumble, the bacterium moves in a new, random direction.
In the presence of an amino acid that is secreted when the organisms are stressed, van Oudenaarden saw that the labeled E. coli swam straight toward the center of a cluster. If they came to the edge, they tumbled. If they found themselves heading into the center of the cluster, toward the attractant, they continued; if not, they tumbled and tried again.
An E. coli cell computes the difference between the attractant concentration now and four seconds ago. In this way, the cell knows if it is going toward or away from the cluster.
"If you could make their memory 10 times longer [than four seconds], their cluster size would be 10 times larger, which would make it much harder for them to stay in contact with each other," van Oudenaarden said. "This makes them more vulnerable. If you could make a genetically altered form of E. coli with a longer memory, that would make it harder for them to form such compact clusters."
The MIT team's web page is at http://web.mit.edu/biophysics.
A version of this article appeared in MIT Tech Talk on October 29, 2003.