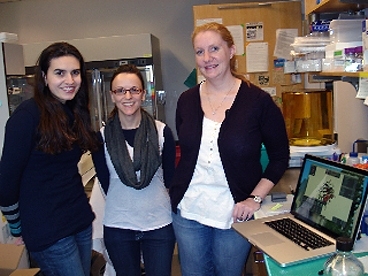
MIT researchers Kelly Knee, Kate Moreau and Oksana Sergeeva — going by the name “Team Crystallin” — came in first among 25 teams in the 2010 University Protein Folding Challenge.
The three members of Biology Professor Jonathan King's lab — Knee and Moreau are postdoctoral fellows and Sergeeva is a graduate student — will share the $5,000 prize, which is awarded to the team that did the best job of using Foldit software to fold an overexpressed protein linked to pancreatic cancer. The three researchers study protein folding, misfolding and chaperones in King’s lab.
The basic idea behind the contest was that by returning a misfolded protein to its optimal structure they would be treating the cancer, at least virtually. Proteins control every process in the human body and are made up of amino acids, whose configuration creates a unique shape for each protein. The shape of the protein determines its function. When folding goes awry and creates the wrong shape, problems and diseases can result. The Foldit software functions like a puzzle that requires the user to apply their problem-solving abilities to predict protein structure.
King said the contest has more significance than may meet the eye.
"How long necklace-like protein chains fold up into their tight and precise three-dimensional folds is a major unsolved problem in modern biology and chemistry," he said. "Computational approaches such as the Foldit competition recruit talented students with a range of backgrounds to the problem. Our team had developed deep insight into protein folding processes through their NIH-funded research on the eye lens crystallins, as well as the ability to work together effectively as a team. We are very proud of their accomplishment."
Moreau also saw the connections between her research and the contest.
“Since we study protein folding in our lab, we thought it would be a fun challenge to apply what we’ve learned at the bench to solving this puzzle,” she said. “We were able to pair our intuition about what the protein conformation should be with the computational models made available by Foldit to make the protein look the way we thought it should.”
Her teammate, Kelly Knee, added, “Foldit is a great tool for fostering interest in science using the protein folding problem as a jumping off point.”
The contest was sponsored by MedImmune, an international pharmaceutical firm that studies how humans can use computers to learn more about proteins and thus better target proteins with drugs for more effective treatment of diseases. The contest was open to undergraduate and graduate students or postdoctoral fellows from top-tier universities in the U.S. and U.K.
The three members of Biology Professor Jonathan King's lab — Knee and Moreau are postdoctoral fellows and Sergeeva is a graduate student — will share the $5,000 prize, which is awarded to the team that did the best job of using Foldit software to fold an overexpressed protein linked to pancreatic cancer. The three researchers study protein folding, misfolding and chaperones in King’s lab.
The basic idea behind the contest was that by returning a misfolded protein to its optimal structure they would be treating the cancer, at least virtually. Proteins control every process in the human body and are made up of amino acids, whose configuration creates a unique shape for each protein. The shape of the protein determines its function. When folding goes awry and creates the wrong shape, problems and diseases can result. The Foldit software functions like a puzzle that requires the user to apply their problem-solving abilities to predict protein structure.
King said the contest has more significance than may meet the eye.
"How long necklace-like protein chains fold up into their tight and precise three-dimensional folds is a major unsolved problem in modern biology and chemistry," he said. "Computational approaches such as the Foldit competition recruit talented students with a range of backgrounds to the problem. Our team had developed deep insight into protein folding processes through their NIH-funded research on the eye lens crystallins, as well as the ability to work together effectively as a team. We are very proud of their accomplishment."
Moreau also saw the connections between her research and the contest.
“Since we study protein folding in our lab, we thought it would be a fun challenge to apply what we’ve learned at the bench to solving this puzzle,” she said. “We were able to pair our intuition about what the protein conformation should be with the computational models made available by Foldit to make the protein look the way we thought it should.”
Her teammate, Kelly Knee, added, “Foldit is a great tool for fostering interest in science using the protein folding problem as a jumping off point.”
The contest was sponsored by MedImmune, an international pharmaceutical firm that studies how humans can use computers to learn more about proteins and thus better target proteins with drugs for more effective treatment of diseases. The contest was open to undergraduate and graduate students or postdoctoral fellows from top-tier universities in the U.S. and U.K.