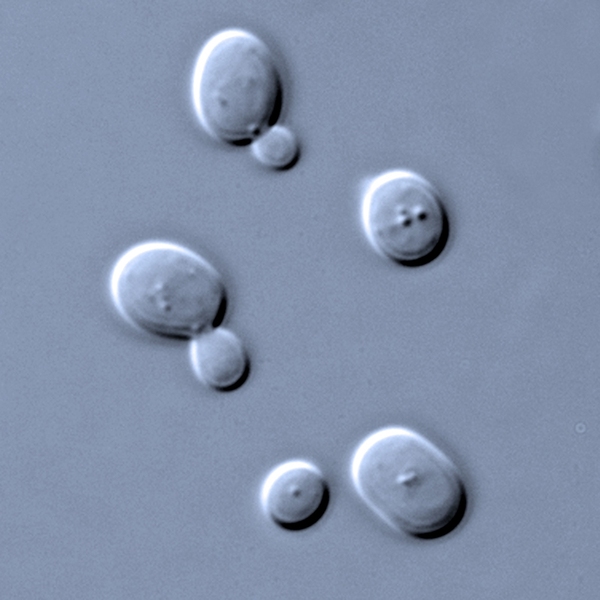
Drug companies often use yeast to manufacture drugs, especially proteins such as antibodies and enzymes. It has been assumed that a batch of genetically identical yeast will secrete such drugs at uniform rates, but MIT chemical engineers have made the surprising discovery that drug productivity varies greatly among individual yeast cells.
The research team, led by J. Christopher Love, assistant professor of chemical engineering, found that while a small subset of yeast is highly productive, a significant minority of the population releases nothing at all. The finding, reported in the journal Biotechnology and Bioengineering, could give drug companies new targets to maximize yeast productivity.
“This is clearly a very powerful tool for selecting cells based on their rate of synthesis, which has a number of applications, including the obvious ability to pick out high-producing cell lines,” says Barry Buckland, a former research and development executive at Merck Research Laboratories, who was not involved in the research.
Love and his colleagues studied a strain of yeast known as Pichia pastoris, which can be used to manufacture proteins including antibodies, growth factors, enzymes and erythropoietin (a hormone that controls red blood cell production). There are now 151 such proteins approved for therapeutic use in the United States or Europe. In 2008, sales of these biopharmaceuticals in the United States exceeded $45 billion, with nearly half of them produced by microbes, including yeast.
To measure yeast productivity, the researchers used a technique that Love had previously developed and used to study immune cells, specifically B cells, called microengraving. The technique allows researchers to look at the quantities of proteins released by single cells.
To do this, they place cells into individual wells arranged in a lattice pattern on a soft rubber surface. Borrowing from an artistic engraving technique used for printmaking, the researchers use that array of cells to “print” the proteins produced by the cells onto the surface of multiple identical glass slides. By measuring how much protein each cell “prints” onto the slides, the researchers can determine the productivity of individual cells.
The yeast in the study were engineered to produce a fragment of a human antibody molecule, but the researchers were surprised to find that a subset of the yeast population (about 35 percent) did not secrete measureable amounts of protein.
“At any given time, there’s a large fraction of the culture that does not contribute to the overall yield,” says Kerry Routenberg Love, a postdoctoral associate in chemical engineering and lead author of the paper.
Intrigued, the researchers ran a longer study, over an eight-hour period, and identified three subpopulations among the cells: those that secrete very little protein over this time, those that secrete consistently high quantities, and those that fluctuate between states of high and low productivity.
Because the yeast used in the study (and in drug manufacturing) are genetically identical, genetic differences cannot account for the discrepancies in productivity. Instead, the difference appears to be epigenetic — meaning that it involves differences in the processing that the proteins undergo after they are assembled, such as protein folding and secretion from the cell.
The researchers are not sure yet why these differences occur, but they have some theories. “It could be due to stress imposed on the cells that have been engineered to overproduce a product that’s not a natural gene product for them,” says Routenberg Love.
Even though the protein itself is not toxic to the cells, it can have adverse effects such as overloading the cell’s protein-folding machinery and secretory pathways. When the cells get overloaded, they need to stop and “reset.”
In future work, the researchers hope to discover more about why there is such variability in yeast productivity, which could lead to new ways to improve drug yields by enhancing productivity in low-producing cells.
The research team, led by J. Christopher Love, assistant professor of chemical engineering, found that while a small subset of yeast is highly productive, a significant minority of the population releases nothing at all. The finding, reported in the journal Biotechnology and Bioengineering, could give drug companies new targets to maximize yeast productivity.
“This is clearly a very powerful tool for selecting cells based on their rate of synthesis, which has a number of applications, including the obvious ability to pick out high-producing cell lines,” says Barry Buckland, a former research and development executive at Merck Research Laboratories, who was not involved in the research.
Love and his colleagues studied a strain of yeast known as Pichia pastoris, which can be used to manufacture proteins including antibodies, growth factors, enzymes and erythropoietin (a hormone that controls red blood cell production). There are now 151 such proteins approved for therapeutic use in the United States or Europe. In 2008, sales of these biopharmaceuticals in the United States exceeded $45 billion, with nearly half of them produced by microbes, including yeast.
To measure yeast productivity, the researchers used a technique that Love had previously developed and used to study immune cells, specifically B cells, called microengraving. The technique allows researchers to look at the quantities of proteins released by single cells.
To do this, they place cells into individual wells arranged in a lattice pattern on a soft rubber surface. Borrowing from an artistic engraving technique used for printmaking, the researchers use that array of cells to “print” the proteins produced by the cells onto the surface of multiple identical glass slides. By measuring how much protein each cell “prints” onto the slides, the researchers can determine the productivity of individual cells.
The yeast in the study were engineered to produce a fragment of a human antibody molecule, but the researchers were surprised to find that a subset of the yeast population (about 35 percent) did not secrete measureable amounts of protein.
“At any given time, there’s a large fraction of the culture that does not contribute to the overall yield,” says Kerry Routenberg Love, a postdoctoral associate in chemical engineering and lead author of the paper.
Intrigued, the researchers ran a longer study, over an eight-hour period, and identified three subpopulations among the cells: those that secrete very little protein over this time, those that secrete consistently high quantities, and those that fluctuate between states of high and low productivity.
Because the yeast used in the study (and in drug manufacturing) are genetically identical, genetic differences cannot account for the discrepancies in productivity. Instead, the difference appears to be epigenetic — meaning that it involves differences in the processing that the proteins undergo after they are assembled, such as protein folding and secretion from the cell.
The researchers are not sure yet why these differences occur, but they have some theories. “It could be due to stress imposed on the cells that have been engineered to overproduce a product that’s not a natural gene product for them,” says Routenberg Love.
Even though the protein itself is not toxic to the cells, it can have adverse effects such as overloading the cell’s protein-folding machinery and secretory pathways. When the cells get overloaded, they need to stop and “reset.”
In future work, the researchers hope to discover more about why there is such variability in yeast productivity, which could lead to new ways to improve drug yields by enhancing productivity in low-producing cells.