MIT researchers have uncovered a critical difference between flu viruses that infect birds and humans, a discovery that could help scientists monitor the evolution of avian flu strains and aid in the development of vaccines against a deadly flu pandemic.
The researchers found that a virus's ability to infect humans depends on whether it can bind to one specific shape of receptor on the surface of human respiratory cells.
"Now that we know what to look for, this could help us not only monitor the bird flu virus, but it can aid in the development of potentially improved therapeutic interventions for both avian and seasonal flu," said Ram Sasisekharan, MIT Underwood Prescott Professor of Biological Engineering and Health Sciences and Technology, and the senior author of a paper on the work that appears in the Jan. 6 issue of Nature Biotechnology.
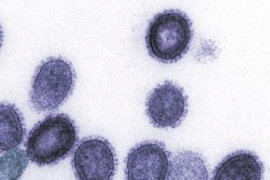
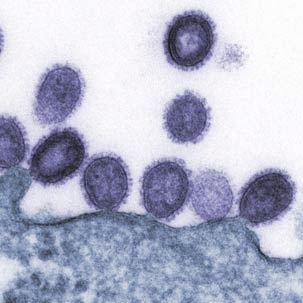
Flu viruses come in many strains, and not all of them can infect humans. Strains known as H1 or H3 have "jumped" from birds to humans and hence are tailored to attack cells of the human upper respiratory tract. H5 strains are usually confined to birds, but when they do infect humans they can have very high fatality rates.
In the past decade, isolated outbreaks of avian flu (H5N1) in humans have raised concerns that a deadly pandemic could arise if the avian flu evolves to a form that can easily infect humans and pass from person to person. Some scientists believe such an outbreak could rival the 1918 "Spanish flu" that killed 50 million to 100 million people worldwide.
Scientists already knew that whether an influenza virus infects humans depends on whether its hemagglutinin, a protein found on the virus surface, can bind to sugar (or glycan) receptors in the respiratory tract. Human respiratory cells have glycan receptors classified as alpha 2-6; avian respiratory cells' glycan receptors are known as alpha 2-3. This classification is based on how the sugars are linked together when they are displayed on cells.
Until now, scientists had believed that a genetic switch that allows the virus to bind to alpha 2-6 receptors instead of alpha 2-3 receptors is responsible for avian viruses' ability to jump to humans.
The MIT study shows that that view does not adequately explain how viruses evolve to infect humans. The new work reveals that, more specifically, it is the ability of a flu virus to bind to a certain shape, or topology, of specific alpha 2-6 glycan receptor that determines whether it will infect humans.
Alpha 2-6 glycan receptors come in two shapes--one that resembles an umbrella, and another that resembles a cone. The MIT team found that to infect humans, flu viruses must bind to the umbrella-shaped alpha 2-6 receptor.
Thus, Sasisekharan and his team have redefined the host receptor for influenza and the criteria for how H5N1 can jump to humans. They did so by showing that the shape of the sugars--and not the type of linkage--is the key determinant for human adaptation of these deadly viruses.
This new interpretation explains inconsistencies that plagued the previous model, according to Sasisekharan. For example, some flu strains that can bind to alpha 2-6 receptors do not infect humans very well. It turns out that those viruses bind to cone-shaped alpha 2-6 receptors, which are present in the human respiratory tract but in much smaller numbers than umbrella-shaped alpha 2-6 receptors.
This new paradigm should help researchers develop a better way to track the evolution of avian flu leading to human adaptation, Sasisekharan said. Now, they know to look for avian viruses that have evolved the ability to bind to umbrella-shaped alpha 2-6 receptors.
That knowledge could help them create vaccines tailored to combat a potential pandemic. Similarly, these findings will help in the development of more effective strategies for seasonal flu, which still is a leading cause of death.
"Subtle changes in influenza viruses over time can dramatically influence the likelihood that these viruses will be able to infect human populations, and this is a huge concern," said Jeremy Berg, director of the National Institute for General Medical Sciences, which funded the research. "This work enables researchers to look at flu viruses in an entirely new way. Dr. Sasisekharan's team achieved this through a multifaceted approach that combines laboratory experiments with the 'mining' of NIH-supported databases, leading to new insights into how the flu virus can adapt to a human host."
Other authors of the Nature Biotechnology paper are Terrence Tumpey of the Centers for Disease Control and Prevention; Aarthi Chandrasekaran, graduate student in MIT's Department of Biological Engineering (BE); Aravind Srinivasan and Karthik Viswanathan, postdoctoral associates in BE; Rahul Raman, research scientist in BE; S. Raguram, visiting scientist in BE; and Viswanathan Sasisekharan, visiting scientist in the Harvard-MIT Division of Health Sciences and Technology.
A version of this article appeared in MIT Tech Talk on January 16, 2008 (download PDF).