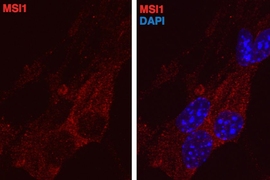
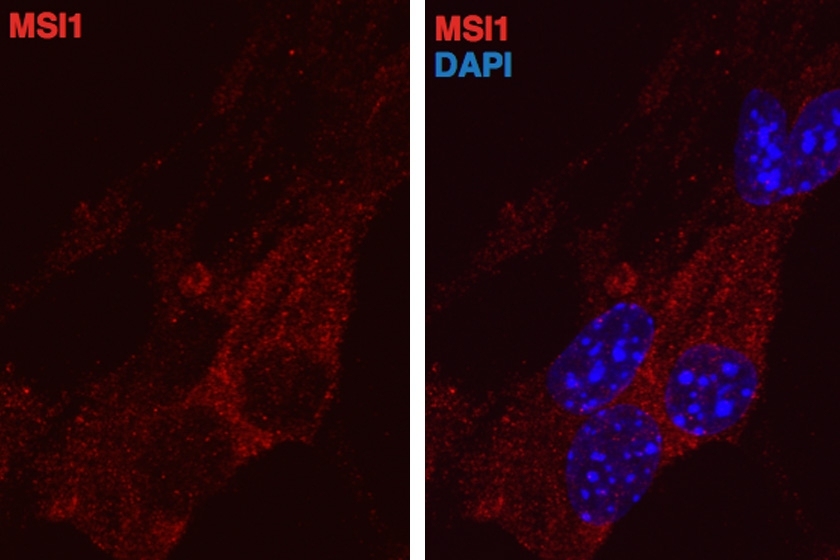
A new study from MIT implicates a family of RNA-binding proteins in the regulation of cancer, particularly in a subtype of breast cancer. These proteins, known as Musashi proteins, can force cells into a state associated with increased proliferation.
Biologists have previously found that this kind of transformation, which often occurs in cancer cells as well as during embryonic development, is controlled by transcription factors — proteins that turn genes on and off. However, the new MIT research reveals that RNA-binding proteins also play an important role. Human cells have about 500 different RNA-binding proteins, which influence gene expression by regulating messenger RNA, the molecule that carries DNA’s instructions to the rest of the cell.
“Recent discoveries show that there’s a lot of RNA-processing that happens in human cells and mammalian cells in general,” says Yarden Katz, a recent MIT PhD recipient and one of the lead authors of the new paper. “RNA is processed at several points within the cell, and this gives opportunities for RNA-binding proteins to regulate RNA at each point. We’re very interested in trying to understand this unexplored class of RNA-binding proteins and how they regulate cell-state transitions.”
Feifei Li of China Agricultural University is also a lead author of the paper, which appears in the journal eLife on Dec. 15. Senior authors of the paper are MIT biology professors Christopher Burge and Rudolf Jaenisch, and Zhengquan Yu of China Agricultural University.
Controlling cell states
Until this study, scientists knew very little about the functions of Musashi proteins. These RNA-binding proteins have traditionally been used to identify neural stem cells, in which they are very abundant. They have also been found in tumors, including in glioblastoma, a very aggressive form of brain cancer.
“Normally they’re marking stem and progenitor cells, but they get turned on in cancers. That was intriguing to us because it suggested they might impose a more undifferentiated state on cancer cells,” Katz says.
To study this possibility, Katz manipulated the levels of Musashi proteins in neural stem cells and measured the effects on other genes. He found that genes affected by Musashi proteins were related to the epithelial-to-mesenchymal transition (EMT), a process by which cells lose their ability to stick together and begin invading other tissues.
EMT has been shown to be important in breast cancer, prompting the team to look into Musashi proteins in cancers of non-neural tissue. They found that Musashi proteins are most highly expressed in a type of breast tumors called luminal B tumors, which are not metastatic but are aggressive and fast-growing.
When the researchers knocked down Musashi proteins in breast cancer cells grown in the lab, the cells were forced out of the epithelial state. Also, if the proteins were artificially boosted in mesenchymal cells, the cells transitioned to an epithelial state. This suggests that Musashi proteins are responsible for maintaining cancer cells in a proliferative, epithelial state.
“These proteins seem to really be regulating this cell-state transition, which we know from other studies is very important, especially in breast cancer,” Katz says.
Embryonic development and cancer
The researchers found that Musashi proteins repress a gene called Jagged1, which in turn regulates the Notch signaling pathway. Notch signaling promotes cell division in neurons during embryonic development and also plays a major role in cancer.
When Jagged1 is repressed, cells are locked in an epithelial state and are much less motile. The researchers found that Musashi proteins also repress Jagged1 during normal mammary-gland development, not just in cancer. When these proteins were overexpressed in normal mammary glands, cells were less able to undergo the type of healthy EMT required for mammary tissue development.
Brenton Graveley, a professor of genetics and developmental biology at the University of Connecticut, says he was surprised to see how much influence Musashi proteins can have by controlling a relatively small number of genes in a cell. “Musashi proteins have been known to be interesting for many years, but until now nobody has really figured out exactly what they’re doing, especially on a genome-wide scale,” he says.
The researchers are now trying to figure out how Musashi proteins, which are normally turned off after embryonic development, get turned back on in cancer cells. “We’ve studied what this protein does, but we know very little about how it’s regulated,” Katz says.
He says it is too early to know if the Musashi proteins might make good targets for cancer drugs, but they could make a good diagnostic marker for what state a cancer cell is in. “It’s more about understanding the cell states of cancer at this stage, and diagnosing them, rather than treating them,” he says.
The research was funded by the National Institutes of Health.